The development of an open and general maternal-fetal physiologically based pharmacokinetic model for drugs metabolized by cytochromes P450 isoenzymes
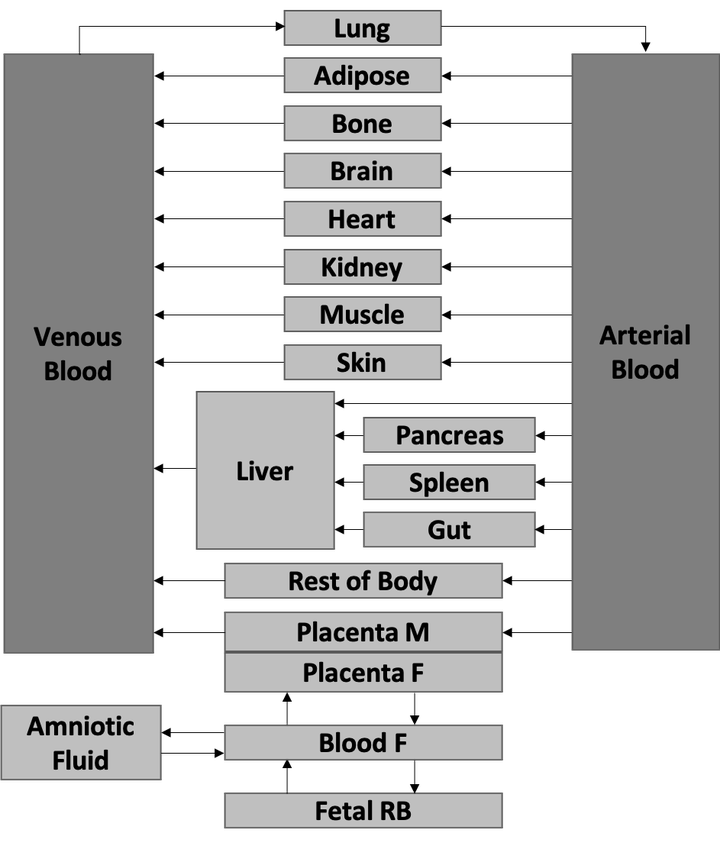 Model structure
Model structurePregnancy causes extensive physiological changes impacting drug exposure in mother and fetus. Predicting a drug’s pharmacokinetic (PK) profile is crucial to ensuring safe and efficacious dosing during pregnancy. Conducting clinical PK trials in pregnancy, however, is both logistically and ethically challenging. Physiologically-based (PB) PK models can provide in silico predictions of drug exposures during pregnancy by accounting for known physiologic changes. These models can guide dosing prior to drug administration and refine dosing once initial exposures are determined.
We developed maternal-fetal and non-pregnant PBPK models implemented as a system of ordinary differential equations in R with mrgsolve to predict maternal/fetal exposure of drugs primarily metabolized by liver CYP450 enzymes (3A4, 2D6, 1A2, 2B6). Model parameters, initially based on literature, were refined using local sensitivity analyses followed by parameter optimization. Models were validated by comparing observed and predicted PK profiles of 10 drugs: midazolam, metoprolol, caffeine, nifedipine, nevirapine, artemether, indinavir, buprenorphine, codeine and methadone.
The relative error (RE) in predicted estimates of area under the curve (AUC) and peak plasma concentration (Cmax) across all tested drugs were 0.17 - 33.1% for AUC and 1.57 - 50.7% for Cmax in the non-pregnant model and 3.34 - 38.1% (AUC) and 7.88 - 23.8% (Cmax) in the pregnant model. Sensitivity analyses and parameter optimization further improved model predictions of these PK parameters. The described PBPK model provides a reproducible, open-source system for model-informed decision for exploring and developing exposure-based dosing recommendations in maternal/fetal patient populations. Inclusion of individual genotype data may further improve predictions by accounting for the difference in clearance between poor and extensive metabolizers.