 Simple Demo of bmediatR in Simulated Data
Simple Demo of bmediatR in Simulated DataMediation analysis focuses on understanding the relationship between an independent variable (X), a dependent variable (Y), and a candidate mediator (M). In a genetic context, X is often a QTL, M a nearby gene, and Y a target of interest. Typical mediation methods use compound hypotheses to test the significance of the pairwise relationship between each variable or require a univariate X and test the indirect effect of X on Y. These methods can be difficult to summarise and are limited by the assumption of a univariate X.
Our approach treats mediation analysis as a model selection problem, exploring the possible relationships between X, M, and Y. Some of these relationships represent mediation, others do not. Each relationship defines a joint likelihood, and does not require univariate X. Combining a prior distribution over the relationships with the joint likelihoods calculated from data provides a posterior distribution over each relationship between X, Y, and M. The method is implemented in the bmediatR R package.
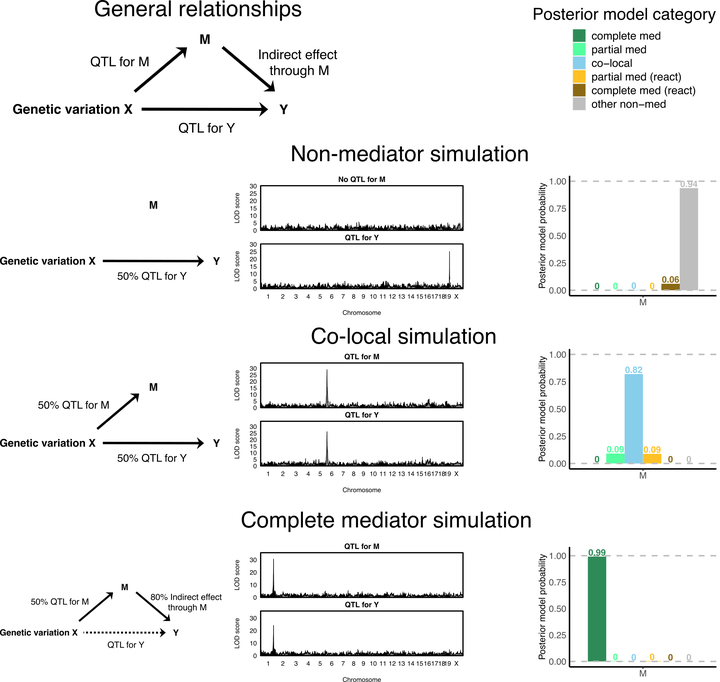
Through simulations we demonstrate the accuracy of bmediatR in inferring the relationship between three variables simulated with varying causal relationships. Comparing these results to existing mediation methods reveals that bmediatR can differentiate a co-local relationship from complete mediation in scenarios where LOD drop cannot. Our examples in data from human cell lines and multiparental mouse populations demonstrate the flexibility of bmediatR in both the inputs and the inferences made.